12. Creating a Dataset from numpy arrays
The goal of this notebook is to demonstrate how to create a xarray.Dataset from scratch. This can be useful if your device is not supported or if you would like to integrate the dtscalibration library in your current routine.
[1]:
import numpy as np
import os
import matplotlib.pyplot as plt
import xarray as xr
from dtscalibration import read_silixa_files
# The following line introduces the .dts accessor for xarray datasets
import dtscalibration # noqa: E401 # noqa: E401
from dtscalibration.variance_stokes import variance_stokes_constant
For a xarray.Dataset object, a few things are needed:
timestamps
Stokes signal
anti-Stokes signal
x (length along fiber)
Let’s grab the data from an existing silixa dataset:
[2]:
filepath = os.path.join("..", "..", "tests", "data", "single_ended")
ds_silixa = read_silixa_files(directory=filepath, silent=True)
We will get all the numpy arrays from this xarray.Dataset to create a new one from ‘scratch’.
Let’s start with the most basic data:
[3]:
x = ds_silixa.x.values
time = ds_silixa.time.values
ST = ds_silixa.st.values
AST = ds_silixa.ast.values
Now this data has to be inserted into an xarray Dataset
[4]:
ds = xr.Dataset()
ds["x"] = ("x", x)
ds["time"] = ("time", time)
ds["st"] = (["x", "time"], ST)
ds["ast"] = (["x", "time"], AST)
[5]:
print(ds)
<xarray.Dataset> Size: 82kB
Dimensions: (x: 1461, time: 3)
Coordinates:
* x (x) float64 12kB -80.74 -80.62 -80.49 -80.36 ... 104.6 104.7 104.8
* time (time) datetime64[ns] 24B 2018-05-04T12:22:17.710000 ... 2018-05...
Data variables:
st (x, time) float64 35kB -0.8058 0.4287 -0.513 ... 27.99 27.83 28.81
ast (x, time) float64 35kB -0.2459 -0.5932 0.1111 ... 36.2 35.7 35.16
For calibration, a few more paramaters are needed:
acquisition time (for calculating residuals for WLS calibration)
reference temperatures
a double ended flag
We’ll put these into the custom xarray.Dataset:
[6]:
ds["acquisitiontimeFW"] = ds_silixa["acquisitiontimeFW"].values
ds["userAcquisitionTimeFW"] = ds_silixa["acquisitiontimeFW"].values
ds["temp1"] = ds_silixa["probe1Temperature"]
ds["temp2"] = ds_silixa["probe2Temperature"]
ds.attrs["isDoubleEnded"] = "0"
Now we can calibrate the data as usual (ordinary least squares in this example).
[7]:
ds = ds.sel(x=slice(-30, 101))
sections = {
"temp1": [slice(20, 25.5)], # warm bath
"temp2": [slice(5.5, 15.5)], # cold bath
}
st_var, resid = variance_stokes_constant(
ds.dts.st, sections, ds.dts.acquisitiontime_fw, reshape_residuals=True
)
ast_var, _ = variance_stokes_constant(
ds.dts.ast, sections, ds.dts.acquisitiontime_fw, reshape_residuals=False
)
out = ds.dts.calibrate_single_ended(sections=sections, st_var=st_var, ast_var=ast_var)
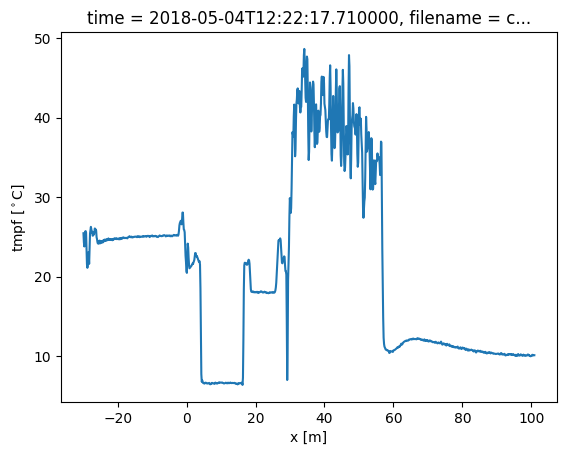
out.isel(time=0).tmpf.plot()
[7]:
[<matplotlib.lines.Line2D at 0x7efbebbf2a00>]